Systems Analysis of Plant Sulfur Metabolism
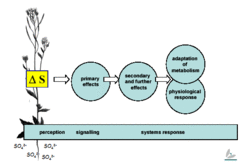
Plants are sessile organisms; if they are to survive and reproduce, they must adapt to the growth conditions in which they find themselves. We use variations in sulfur levels as a stimulus and analyse the complex response using diverse multiparallel techniques, particularly transcript and metabolite profiling, trying to piece together the total system response. The plant of choice here is, obviously, Arabidopsis thaliana, although results obtained in this model system are likely to be transferable to other plant species and crop plants. The goal is to provide a consistent and holistic description of plant sulfur metabolism and its regulation.
H Hesse and R Höfgen (2001) Application of Genomics in Agriculture. In: Molecular analysis of plant adapatation to the environment. Eds: Malcolm J. Hawkesford, Peter Buchner. Kluwer AP, Dordrecht, The Netherlands, 61-79
V Nikiforova, J Freitag, S Kempa, M Adamik, H Hesse, R Hoefgen (2003) Transcriptome analysis of sulfur depletion in Arabidopsis thaliana: Interlacing of biosynthetic pathways provides response specificity. The Plant Journal, 33, 633-650.
Regulation
Plants adapt to available sulfur soil levels by regulating gene expression and protein activity to maintain homeostasis. Sulfur availability in the environment is not static, nor is the plant's dependence on sulfur at various developmental stages. Thus, one can assume not only that the activities of regulatory proteins are dynamic, but also that changes in the expression of transcription factors involved in triggering downstream gene expression change temporally. Sulfur deprivation triggers a slow adaptive process that resets the level of sulfur homeostasis. Using transcript profiling, we have been able to identify a number of transcription factors involved in this process, which are now the target of further investigations.
Metabolome analysis and bioinformatics
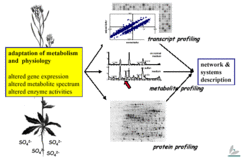
Gene expression, metabolite spectrum and enzyme activities change under sulfur-limiting conditions.
The response of steady state transcription levels to the sulfur stimulus is but the first chapter of the story. To understand the system response, we have to turn the page and look at protein profiles - levels and activities - and before closing the book, at metabolite profiles, which adjust rapidly in response to changes in protein expression. We are now focusing on metabolome analysis: The same samples used for transcriptome analysis are examined using element analysis (ICP-AES) and metabolite analyses (HPLC, CE, GC/MS, GC/TOF, LC/MS), either in house or in collaboration with outside research groups.
Malcolm J. Hawkesford, Rothamsted Research, UK
As these analyses are refined and data accumulates, it will become more and more important to overlay and compare transcript and metabolite profiles in order to try to generate an in silico representation of the plant sulfur regulatory complement. Various approaches are and will be followed here: bioinformatic tools have to be developed and/or adapted to fully mine the data. Otherwise, it will not be possible to fully describe the system: by looking only at the most highly expressed genes in isolation, we would simply be scratching at the surface.
Transcriptome Analysis
Plants and some photoautotrophic bacteria assimilate inorganic sulfur from sulfates into cysteine, the first sulfur-containing organic compound and, effectively, the sole molecular doorway for reduced sulfur in all living beings. This essential process has been as finely tuned through millennia of evolution as photosynthesis. Cysteine is subsequently converted to methionine, and then into a variety of other sulfur-containing organic compounds. Sulfur assimilation is even more spendy in terms of reduction equivalents than nitrogen assimilation. Obviously, such a costly enterprise is highly controlled in juxtaposition with the rest of metabolism.
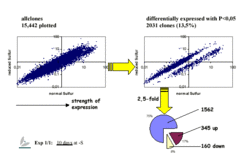
To elucidate this network of interactions, we stimulate Arabidopsis with sulfur (i.e. sulfate) at its rhizosphere with various concentrations and at different developmental stages to institute periods of starvation and replenishment. The plant tissue samples (roots, shoots) are then subjected to array hybridisation/transcript profiling after RNA extraction using either macro-arrays of 7,200 non-redundant genes on nylon filters and now full genome chips. The expression profiles are processed to select differentially expressed genes. Depending on the duration of treatment, anything between a handful and thousands of genes exhibit altered expression mirroring the gradual response of the system to conditions of altered sulfur availability. Among these responsive genes we expect to find sulfur-regulated genes; genes involved in perception, signalling, and immediate responses; and genes further down the line involved in more pleiotropic mechanisms like general stress responses. Since they arise in response to sulfur stimulation, the latter are still regarded as sulfur-responsive genes.
Sulfur-responsive genes are grouped by functional category or biosynthetic pathway. As expected, genes of the sulfur assimilation pathway are altered in expression. Furthermore, genes involved in the flavonoid, auxin, and jasmonate biosynthesis pathways are up regulated when sulfur is limiting. We focus most of our attention, however, on the regulatory elements, transcription factors.
V Nikiforova, J Freitag, S Kempa, M Adamik, H Hesse, R Hoefgen (2003) Transcriptome analysis of sulfur depletion in Arabidopsis thaliana: Interlacing of biosynthetic pathways provides response specificity. The Plant Journal, 33, 633-650
Further reading
MY Hirai, T Fujiwara, M Awazuhara, T Kimura, M Noji, K Saito (2003) Global expression profiling of sulfur-starved Arabidopsis by DNA macroarray reveals the role of O-acetyl-L-serine as a general regulator of gene expression in response to sulfur nutrition. Plant Journal. 33(4)651-663
A Maruyama-Nakashita, E Inoue, A Watanabe-Takahashi, T Yarnaya, and H Takahashi (2003) Transcriptome profiling of sulfur-responsive genes in Arabidopsis reveals global effects of sulfur nutrition on multiple metabolic pathways. Plant Physiology. 132(2)597-605
Sulfur and Other Plant Nutrients
The plant sulfur assimilation pathway is intricately interconnected with various other pathways and regulatory circuits. This nutritional wiring scheme is the next level to be investigated. We will approach this mainly through co-operations in house with the groups of Michael Udvardi, Georg Leggewie, Wolf-Ruediger Scheible, Mark Stitt, Joachim Kopka, and Oliver Fiehn either as joint experimentation or the mutual use of databases. In addition, external collaborations are in place for various aspects of this work and we are open to expanding this network further.


