Viral Replication and Plant Tolerance
Viruses can cause extreme change in the cellular environment of their host. Our goal is to understand what are the conserved mechanisms deployed by plant viruses to shape a cellular environment conductive to replication of their genetic material and the matching plant pathways that protect host cell against virus-induced damage.
Viruses hijack host membranes
RNA viruses form replication complexes in different organelles. These proviral structures segregate viral factors and co-opted host components from the cytoplasm. Many studies in both plants and animals have shown the extent to which viruses remodel host membrane during the establishment of their viral factories, and how ubiquitous this replicative strategy is, hinting at a deeply conserved feature of the viral life cycle. Despite their apparent importance, little is known about how these structures are formed. How did viral RNA replication evolve to be able to so fundamentally change organelle morphology?
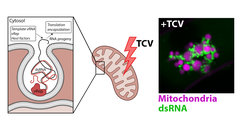
Right panel: VRCs captured in live Arabidopsis roots infected with TCV captured by confocal microscopy. Mitochondria are dyed in magenta and dsRNA is visualized with GFP fluorescence in green.
adapted from: Clavel et Dagdas 2021 - Current Opinion in Plant Biology, 63:102106
To answer this question we employ proteomics, biochemistry, cell biology and genetics, to understand how key viral proteins orchestrate organelle morphology. We undertake a comparative mechanistic approach in different species that span plant diversity in order to gain an evolutionary perspective as to how these structures came to be. Our research has the potential to identify previously unknown aspects of the viral life cycle that could be exploited for growing crops resilient to infection.
Our key research questions include (but are not limited to):
- What are the conserved host factors necessary and sufficient to assemble a replication organelle?
- How do viral factors orchestrate the assembly of a replication organelle?
- Do different viruses assemble different replication complexes? Conversely, do distinct organelles result in distinct replication complexes?
Plants respond and cope with viral-induced organelle damage

We recently discovered a mechanism employed by plants to deal with damage caused by viral hijacking of their organelle membranes. When infected by different viruses, Arabidopsis plants deploy a novel selective autophagy pathway that degrades toxic material otherwise accumulating in the cytosol. Accordingly in absence of functional autophagy, infected plants display much more severe symptoms and develop spontaneous dead lesions. Thus, we have uncovered an unexpected pro-life function of selective autophagy and wish to understand its molecular intricacies. To achieve this, we focus on newly identified molecular players that define this pathway and bridge the gap between toxic cargo and the autophagy machinery.
We are most curious about:
- How do these novel proteins switch from their normal function to their autophagy-related function?
- What defines the toxicity of the autophagy cargo during infection?
- What signal(s) is/are necessary to kickstart the autophagy process?
- Can we identify triggers outside infection that rely on this pathway?
- What defines the early, cell-autonomous response of plant cells when viral replication is perceived?

