The Genetics of Crop Metabolism
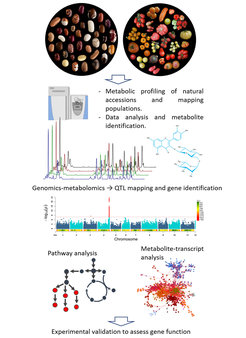
The plant kingdom is capable of producing enormous structurally and functionally diverse compounds. Some of these compounds are essential for plant growth and development such amino acids, sugars and organic acidy. However, the vast majority are secondary metabolites (also called ‘specialized metabolites’) which have important roles in the interaction between the plant and its environment as well as being unique sources for human health benefits. Our lab uses metabolomics techniques (both LC-MS and GC-MS) to evaluate the metabolic compositions across a wide range of plant species. We use Genome-Wide Association Studies (GWAS) and mapping populations to identify genes involved in the regulation of secondary metabolites, with particular focus on factors involved in flavonoid, anthocyanin and alkaloid pathways in tomato and common bean.
In addition, the lab is interested in the regulation of plant metabolism associated with stress conditions and fruit and seed quality.
Selected publications:
- Alseekh S, Ofner I, Liu Z, Osorio S, Vallarino J, Last RL, Zamir D, Tohge T, Fernie A.R. (2020). Quantitative trait loci analysis (QTL) of seed specialized metabolites reveals seed-specific flavonols and differential regulation of glycoalkaloid content in tomato. Plant Journal.
- Alseekh, S., Tong, H., Scossa, F., Brotman, Y., Vigroux, F., Tohge, T., Ofner, I., Zamir, D., Nikoloski, Z. and Fernie, A.R. (2017) Canalization of Tomato Fruit Metabolism. The Plant Cell, 29, 2753-2765.
- Alseekh, S., Tohge, T., Wendenberg, R., Scossa, F., Omranian, N., Li, J., Kleessen, S., Giavalisco, P., Pleban, T., Mueller-Roeber, B., Zamir, D., Nikoloski, Z. and Fernie, A.R. (2015) Identification and mode of inheritance of quantitative trait loci for secondary metabolite abundance in tomato. The Plant cell, 27, 485-512.
- Garbowicz, K., Liu, Z., Alseekh, S.*, Tieman, D., Taylor, M., Kuhalskaya, A., Ofner, I., Zamir, D., Klee, H.J., Fernie, A.R. and Brotman, Y. (2018) Quantitative Trait Loci Analysis Identifies a Prominent Gene Involved in the Production of Fatty Acid-Derived Flavor Volatiles in Tomato. Molecular plant, 11, 1147-1165.
- Zhang Y, Butelli E, Alseekh S, Tohge T, Rallapalli G, Luo J, Kawar P, Hill L, Santino A, Fernie AR, Cathie Martin C (2015). Multi-level Engineering Facilitates the Production of Phenylpropanoid Compounds in Tomato. Nature Commun. 6-8635



