Pathway Analysis of Sulfur Containing Amino Acids
Pathway Regulation
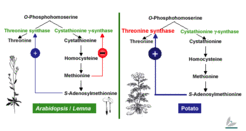
Sulfate metabolism in plants is regulated through a complex circuit that is itself affected by multiple enzymes with various subcellular locations and by differences in the developmental and spatial activity patterns of these enzymes. Molecular engineering of sulfate assimilation serves as a tool for investigating pathways at molecular, biochemical, and physiological levels: Only an in-depth fundamental understanding of these pathways will enable the rational and targeted approaches required for successful plant breeding.
In order to gain insight into the control mechanisms involved in sulfur-containing amino acid biosynthesis, we are analysing the expression levels of involved genes by isolating and studying the promoters of key enzymes. In addition, spatial and developmental aspects of regulation are investigated with respect to enzyme expression and activity at both tissue and subcellular levels. The roles of catabolism and recycling mechanisms in this regulation will be points of interest in the future.
H Hesse and R Hoefgen (2003) Molecular aspects of methionine biosynthesis. TIPS, 8, 259-262
O Kreft, R Hoefgen, and H Hesse (2003) Functional Analysis of Cystathionine gamma-Synthase in Genetically Engineered Potato Plants. Plant Physiology, 131, 1843-1854
V Nikiforova, J Freitag, S Kempa, M Adamik, H Hesse, R Hoefgen (2003) Transcriptome analysis of sulfur depletion in Arabidopsis thaliana: Interlacing of biosynthetic pathways provides response specificity. The Plant Journal, 33, 633-650
V Nikiforova, S Kempa, M Zeh, S Maimann, O Kreft, A P Casazza, K Riedel, E Tauberger, R Hoefgen, H Hesse (2002) Engineering of cysteine and methionine biosynthesis in potato. Amino Acids, 22, 259-278
